Fields of research
ICIPS - FOR5098
“Innovation and Coevolution in Plant Sexual Reproduction”
ICIPS is a DFG-funded research group coordinated by Prof. Dr. Annette Becker in Gießen

Funding: 2021-2025 German Research Foundation (DFG)
Coordination: Romain.Scalone@bot.jlug.de
How do evolutionary novelties arise?
One of the central goals of evolutionary research is to understand how changes in gene regulatory networks leads to the origin of novel, complex traits, or, in short: How do evolutionary novelties arise? We propose to use the origin of novel traits emerging in the order Ranunculales (buttercups) as model system to study the molecular nature of their origin. Ranunculales are early branching eudicots that comprise morphologically diverse species such as buttercups, poppies, columbines and larkspur. Ranunculales are special because several morphological novelties evolved repeatedly, e.g., spurred floral organs and zygomorphy, or the reduction of perianth. Moreover, novel floral organs, reduction to spiral floral phyllotaxy from whorled, dioecy, and wind pollination originated in this order, in some cases repeatedly. Many species within the Ranunculales are amenable to Virus-Induced Gene Silencing, which allows for functional characterization of candidate genes in later stages of the proposed project.
We propose to sequence 20 transcriptomes of eight Ranunculales species (Eschscholzia californica, Papaver somniferum, Capnoides sempervirens, Pteridophyllum racemosum, Thalictrum thalictroides, Aquilegia coerulea, Nigella damascena and Staphisagria picta) that encompass these novelties including two outgroup species. For the species without genome information available, we plan to sequence their genomes. The transcriptomes will be obtained from a variety of tissues, including floral buds at different developmental stages, dissected floral organs, petal time-series and vegetative tissues. Our ultimate goal is to uncover the GRN modules required for the emergence of novel, often convergently emerging traits. Furthermore, we hope to unravel in the future the processes by which GRN modules are reused and coopted to generate complex floral morphologies and thus drive species diversification. In the proposed project, we want to provide the resources for comparative analysis of genomes and transcriptomes in Ranunculales. This will pave the way to unravel the genetic base novel morphological trait origin.
Funding: 2021 - 2025 German Research Foundation (DFG)
Kooperationspartner: Alexander Goesmann (JLU), Elena Kramer (Harvard), Veronica di Stilio (Seattle), Paula Elomaa (Helsinki), Sophie Nadot (Paris), Catherine Damerval (Paris), Florian Jabbour (Paris), Ian Grahan (York).
Contact: Annette.Becker@bot1.bio.uni-giessen.de
The evolution of plasmodesmata

Postcytokinetic formation of secondary PD in pre-existing cell walls is an appropriate mode to adjust PD numbers (and transport capacities) to changing requirements and has evolved (independently?) at least in some lycophyte lineages and spermatophytes. Whether secondary PD formation also occurs in other land plant lineages is unclear and will be investigated in the project.
Since the project is part of the DFG priority programme 2237 “MAdLand - Molecular Adaptation to Land: plant evolution to change”, we will perform our microscopic analyses on the MAdLand model plant species which represent distinct taxa of streptophyte algae and non-seed land plants. We aim to discover basic differences in PD structure and origin among the streptophyte lineages, the molecular basis of which will be addressed in the second funding period.
Funding: 2020-2023 German research Foundation (DFG)
Contact: Katrin.Ehlers@bot1.bio.uni-giessen.de
Cooperation partners: Stefan Rensing, University of Marburg (MAdLand coordinator), cf. https://madland.science/ and http://madland.science/projects.php
Phylotranscriptomics of the carpel developmental toolkit - an evodevo study towards understanding the origin of flowering plants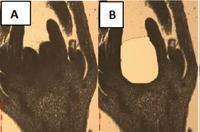
We know relatively little about the origin of angiosperms, a group of plants that dominate most terrestrial ecosystems and provides us with most of our food. Despite the fact that genomes and transcriptomes of angiosperms and their sister group, the gymnosperms are available already; some early angiosperm fossils are described; and the phylogeny of seed plants, comprising angiosperms and seed plants is generally solved, answers to the key questions of how did the first angiosperm look like and which genes were required to build this ancestral angiosperm remain unanswered.
The general aim of this project is to identify a minimal set of genes required for carpel development in angiosperms to understand which genetic prerequisites were required to build the ancestral carpel, and thus, how a crucial step during angiosperm origin was completed. Subsequently, the molecular functions and genetic interactions of this set of genes will be identified to predict the ancestral state of the carpel development network and to learn which genes carried out which functions in the ancestral angiosperm. This project is the first to systematically analyze carpel developmental regulators in an unbiased way across a wide range of phylogenetically important taxa by a combination of laser capture microdissection and phylotranscriptomics. With the newly published genomes of A. trichopoda and P. abies and the availability of the W. mirabilis transcriptomes as second gymnosperm from a divergent lineage, this project is now a timely and realistic undertaking.
Funding: 2016 – 2019 German Research Foundation (DFG)
Cooperation partners: Alexander Goesmann and Oliver Rupp, Institute for Bioinformatics and Systems Biology, JLU Gießen, Knut Beuerlein, Rudolph-Buchheim-Institute for Pharmacology, JLU Gießen
Contact: Annette.Becker@bot1.bio.uni-giessen.de
Evolutionary genetics of carpel development using California poppy (Eschscholzia californica) as a new model species

Funding: 2005-2014 German Research Foundation (DFG), continuation with JLU funding
Contact: Annette.Becker@bot1.bio.uni-giessen.de
