Prof. Dr. R. Renkawitz
 R. Renkawitz |
Prof Dr. R. Renkawitz
Institute for Genetics Justus-Liebig-University Gießen Heinrich-Buff-Ring 58-62 35392 Gießen Tel: +49 (0) 641-99-35486 Fax: +49 (0) 641-99-35469 E-Mail: rainer.renkawitz |
Research
Epigenetic Mechanisms Controlling Gene Activity
A large fraction of the 25 000 genes in the mammalian genome is repressed in many cell types and tissues. Within a cellular population most of the repressed genes stay inactive, thus the epigenetic status is transferred to the daugther cells. The maintenance of the repressed state is a fundamental requirement for embryogenesis, cell differentiation and tissue function. Genome-wide epigenetic DNA and chromatin modification play essential roles in regulating gene expression.
An altered pattern of epigenetic modifications is involved in many common human diseases, including cancer, Tumor-suppressor genes are often turned off, whereas proto-oncogenes are activated. Therefore, carcinogenesis can result from hypermethylation and from hypomethylation of genomic regions. To date, for many human genetic diseases, the underlying genetic defect has been determined. Nevertheless, the open and important question remains unanswered, how gene repression is mediated and maintained. The molecular mechanisms involved include chromatin remodeling, histone modification and DNA methylation. The molecular function of these in gene repression, gene activation, gene insulation and subnuclear arrangement are studied in detail in our group.
Advances in understanding epigenetic mechanisms have been and will be accompanied by new therapeutic options and targets for treatment.
Research Focus:
|
|
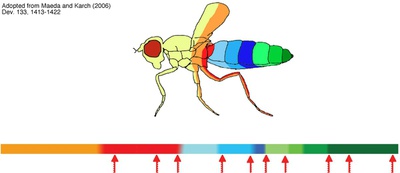
Key mediators of the enhancer blocker function of Drosophila CTCF
Functional eukaryotic gene units including all of their regulatory elements are insulated from each other to allow the action of regulatory elements on the appropriate gene. We and others have analyzed the DNA binding factor CTCF, which is the only protein known to mediate a directional blocking activity of enhancer elements in vertebrates (3, 8). In focusing on Drosophila CTCF, we found out that dCTCF mediates enhancer blocking on the Fab-8 insulator and binds to several other known insulator sequences (1, 2, 9). One set of dCTCF binding sites separates the homeotic genes responsible for the segmental identity in flies (2, 5). Deletion of dCTCF causes a defect in abdominal segmentation (homeotic mutation) and lethality (1). Search for co-factors involved in dCTCF function revealed a new, unknown protein (7), se veral proteins known to mediate chromatin remodelling (4, 5, 6) and chromatin modification (10). These results support a model for insulator function: DNA bound dCTCF recruites several co-factors removing changes in chromatin modification, which may have spread from flanking regions. |
|
1. Mohan, M., Bartkuhn, M., Herold, M., Philippen, A., Heinl, N., Bardenhagen, I., Leers, L., White, R.A.H., Renkawitz-Pohl, R., Saumweber, H. and Renkawitz, R. (2007). The Drosophila insulator proteins CTCF and CP190 link enhancer blocking to body patterning. EMBO J 26, 4203-4214. 2. Bartkuhn, M., Straub, T., Herold, M., Herrmann, M., Rathke, C., Saumweber, H., Gilfillan, G.D., Becker, P.B. and Renkawitz, R. (2009). Active promoters and insulators are marked by the centrosomal protein 190. EMBO J. 28, 877-888. 3. Herold, M., Bartkuhn, M. and Renkawitz, R. (2012) CTCF: Insights into insulator function during development. Development 139, 1045-1057. 4. Ahanger, S.H., Günther, K., Weth, O., Bartkuhn, M., Bhonde, R.R., Shouche, Y.S., and Renkawitz, R. (2014). Ectopically tethered CP190 induces large-scale chromatin decondensation. Scientific Reports 4, 3917. 5. Bohla, D.*, Herold, M*, Panzer, I, Buxa, M.K., Ali, T., Demmers, J., Krüger, M., Scharfe, M., Jarek, M., Bartkuhn, M.1, Renkawitz, R. (2014). A functional insulator screen identifies NURF and dREAM components to be required for enhancer-blocking. PLoS One 9: e107765. 6. Weth, O., Paprotka, C., Günther, K., Schulte, A., Baierl, M., Leers, J., Galjart, N. and Renkawitz, R. (2014). CTCF Induces Histone Variant Incorporation, Erases the H3K27me3 Histone Mark and Opens Chromatin. Nucl. Acids Res. 42, 11941-11951. 7. Maksimenko, O.*, Bartkuhn, M.*, Stakhov, V.*, Herold, M.*, Zolotarev, N., Jox, T., Buxa, M.K., Kirsch, R., Bonchuk, A., Fedotova, A., Kyrchanova, O., Renkawitz, R., Georgiev, P. (2015) Two new insulator proteins, Pita and ZIPIC, target CP190 to chromatin. Genome Res. 25, 89-99. 8. Ali, T., Renkawitz, R. and Bartkuhn, M. (2016). Insulators and domains of gene expression. Curr Opin Genet Dev 37, 17-26. 9. Buxa, M.K., Slotman, J.A., van Royen, M.E., Paul, M.W., Houtsmuller, A.B. and Renkawitz, R. (2016). Insulator speckles associated with long-distance chromatin contacts. Biology Open 5, 1266-1274. 10. Ali, T., Krüger, M., Bhuju, S., Jarek, M., Bartkuhn, M. and Renkawitz, R. (2017). Chromatin binding of Gcn5 in Drosophila is largely mediated by CP190. Nucl. Acids Res. 45, 2384-2395. |
|
|
|
Role of multi zinc finger proteins in the dynamic change of the nuclear architecture during cell cycle and differentiation
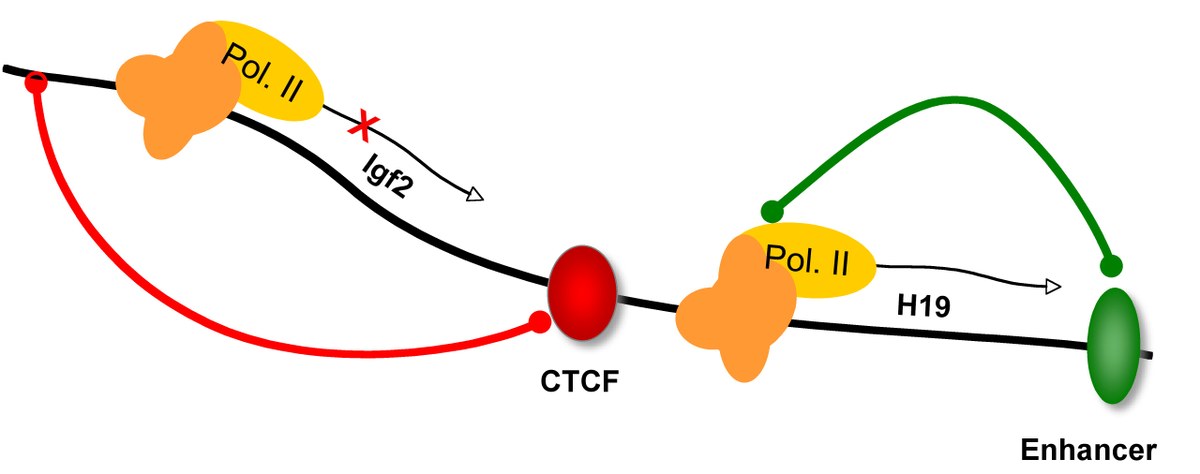
The multi-zinc finger proteins CTCF and CTCFL are unique and conserved factors with a role in transcriptional regulation, the organization of chromatin into distinct domains and imprinting (2, 4). CTCF is ubiquitously expressed with cell-specific and gene-specific functions (3, 5, 6, 7). In contrast, the related factor CTCFL is expressed in the testis (8). Abnormal up-regulation of CTCFL, on the other hand, may be linked to tumorigenesis. Thus, while binding to similar sites in the genome, these proteins do have distinct roles. We could show that CTCF, in contrast to CTCFL and to other zinc finger proteins, remains bound to mitotic chromatin (1). Despite very similar zinc finger domains of both proteins, they differ in their molecular function. Specificity of chromatin binding is driven by nucleosomal modification and density (8). CTCF is actively opening compact chromatin, whereas CTCFL cannot (9). |
|
1. Burke, LJ., Zhang, R., Bartkuhn, M., Tiwari, VK., Tavoosidana, G., Kurukuti, S., Weth, C., Leers, J., Galjart, N., Ohlsson, R., and Renkawitz, R. (2005) CTCF binding and higher order chromatin structure of the H19 locus are maintained in mitotic chromatin. EMBO J. 24, 3291-3300. 2. Bartkuhn, M. and Renkawitz, R. (2008) Long range chromatin interactions involved in gene regulation. Biochim. Biophys. Acta 1783, 2161-2166. 3. Heath, H., de Almeida, C.R., Sleutels, F., Dingjan, G., van de Nobelen, S., Jonkers, I., Ling, K.W., Gribnau, J., Renkawitz, R., Grosveld, F., Hendriks, R.W. and Galjart, N. (2008). CTCF regulates cell cycle progression of αβ T cells in the thymus. EMBO J. 27, 2839-2850. 4. Ohlsson, R., Bartkuhn, M. and Renkawitz, R. (2010). CTCF shapes chromatin by multiple mechanisms. Chromosoma 119, 351-360. 5. Weth, O., Weth, C., Bartkuhn, M., Leers, J., Uhle, F. and Renkawitz, R. (2010) Modular Insulators: Genome wide search for composite CTCF / Thyroid hormone receptor binding sites. PLoS ONE 5, e10119. 6. van de Nobelen, S., Rosa-Garrido, M., Leers, J., Heath1, H., Soochit, W., Joosen, L., Jonkers, I., Demmers, J., van der Reijden, M., Torrano, V., Grosveld, F., Delgado, M.D., Renkawitz, R., Galjart, N. and Sleutels F. (2010) CTCF regulates the local epigenetic state of ribosomal DNA repeats. Epigenetics & Chromatin 3, 19. 7. Weth, O. and Renkawitz, R. (2011) CTCF function is modulated by neighboring DNA binding factors. Biochem. Cell Biol. 89, 459-468. 8. Sleutels, F., W. Soochit, M. Bartkuhn, H. Heath, S. Dienstbach, P. Bergmaier, V. Franke, M. Rosa-Garrido, S. van de Nobelen, L. Caesar, M. van der Reijden, J. C. Bryne, W. van Ijcken, J. A. Grootegoed, M. D. Delgado, B. Lenhard, R. Renkawitz, F. Grosveld and N. Galjart (2012). The male germ cell gene regulator CTCFL is functionally different from CTCF and binds CTCF-like consensus sites in a nucleosome composition-dependent manner. Epigenetics & Chromatin 5(1): 8. 9. Weth, O., Paprotka, C., Günther, K., Schulte, A., Baierl, M., Leers, J., Galjart, N. and Renkawitz, R. (2014). CTCF Induces Histone Variant Incorporation, Erases the H3K27me3 Histone Mark and Opens Chromatin. Nucl. Acids Res. 42, 11941-11951
|
|