PhD projects
Collection of currently running PhD projects.
PhD project of Julian Hahnfeld: Exploring Small Proteins: Advancing Prediction with Deep Learning for Bacterial sORFs and Specialized Databases
Small proteins with fewer than 100 and, in particular, fewer than 50 amino acids are still largely unexplored. They are encoded by small open reading frames (sORFs) and represent an important part of the genetic repertoire of bacteria that often remains neglected. In recent years, the development of ribosome profiling protocols has led to an increasing number of newly detected small proteins. Despite this, they are frequently overlooked during computational gene prediction and automated genome annotation. In addition, functional descriptions often cannot be assigned to predicted small proteins due to a lack of homologs with high sequence similarity in public databases. For this reason, new approaches for the in silico prediction of bacterial sORFs and small proteins, as well as specialized small protein databases are needed.
For the prediction approach, deep learning techniques are promising as they have provided excellent results for many conventional biological questions and are capable of self-learning relevant features from sequence data.
Since the gene and protein features of sORFs partly differ from those of longer genes, traditional gene prediction algorithms exhibit only poor prediction performance for sORFs, due to high false positive rates. These features can be used to develop a new sORF specific prediction approach. For this purpose, the current state of known sORFs and small proteins in public databases was investigated to find suitable features and potential biases. Promising features were compared between sORFs and long ORFs, such as the relative GC content of genes, amino acid composition, transcription initiation and termination mechanisms, and physicochemical properties of proteins.
Based on these distinct features, a new deep-learning-based approach for the prediction of sORFs will be developed and analyzed in terms of feature importance and model performance.
PhD project of Linda Fenske: Streptococcus agalactiae: A potential zoonotic pathogen for humans and cattle
Streptococcus agalactiae, from the streptococcus family, is one of the main triggers of mastitis in cattle as well as occasionally in other mammals. However, S. agalactiae also plays an important role as a human infectious agent, as the main cause of neonatal infections, triggered by transmission from mother to newborn, which can result in pneumonia, meningitis or septicemia.
Recent findings also warn of food borne infections. For example, in Singapore in 2015, there was a cluster of severe infections triggered by the consumption of raw fish. So far, it has not been fully clarified whether this is an original transmission from fish to humans or vice versa. It also remains to be clarified whether transmission of the pathogen from cows to humans or vice versa, for example, during the milking process, is possible and, if so, capable of causing a severe infection.
The occurrence of the same sequence types in different host species definitely supports this, but could as well indicate a common source of infection in the environment.
To elucidate how similar strains of bovine and human origin really are a broad comparative genomic analysis of isolates of different origin is performed. Previous studies have mostly focused on non-holistic methods, usually sequencing only those regions of the genome that are of particular interest. In contrast, a bioinformatic analysis on genome level will be conducted. For this purpose, isolates from cattle affected with mastitis as well as clinical human are compared. The main focus is to address the question of zoonotic potential and to determine how host-specific different strains in fact are.
PhD project of Michael Schwabe: The transcriptome of Tineola bisselliella
Keratin is a structural fibrous protein and is the main building block in hair, wool, feathers, horn, and nails. In slaughterhouses and poultry farms, large amounts of keratin-containing waste are generated every year, e.g. in the form of feathers. Keratin is very resistant to physical influences and chemical and biological agents. As a result, it is often buried or burned in landfills. Nevertheless, keratin-containing waste does not accumulate in nature.
Only few microorganisms can break down keratin, and even fewer higher eukaryotes have this ability. One of them is Tineola bisselliella - the common or webbing clothes moth. The mechanism of keratin digestion in beetles, moths and microorganisms is different from one another and the keratin degrading mechanism in the larvae has not yet been fully described. Therefore, we compare the transcriptomic shift in the intestine of T. bisselliella larvae fed with feathers (keratin-rich) and insect carcasses (keratin-free). We search for known and new enzymes that are believed to be part of the keratin degrading system. Our data include potential symbiotic transcripts as well as host transcripts.
The project is initiated and financed by the Fraunhofer IME in Giessen.
PhD project of Patrick Barth
Non-coding RNAs (ncRNAs) are RNA molecules that are not translated into proteins. Nonetheless, they still partake in a variety of essential biological processes and also take important roles in the complex regulatory system of gene expression. In the last decade several new ncRNA classes have been characterized showing that ncRNAs are a current topic with constant discoveries.
To gain more insight into the complex field of ncRNAs the RTG 2355 was arranged consisting of twelve project groups from different disciplines. This project is part of the RTG 2355 and aims at supporting the other members with data analyses and implementing automated workflows which will be made accessible in an easy to use manner. Consequently, collaborations between the members, spanning the different disciplines, are encouraged.
As part of those collaborations an iCLIP-analysis pipeline has been developed and is still further extended by varying postprocessing analyses.
Additionally, a project in which the packaging of siRNAs into exosomes in plants and their potential cross-kingdom gene silencing is being investigated.
PhD project of Andreas Hoek
WASP: A versatile, web-accessible single cell RNA-Seq processing platform
Since its first application in 2009, single cell RNA sequencing (scRNA-seq) has experienced a steep development. Due to the unprecedented resolution of single cell technology it is widely applicable in many different fields of research, ranging from basic research questions such as analyzing differentiation processes in cells up to highly specific biomedical questions such as tumor cell characterisation. Furthermore, scRNA-seq has undergone many developments in the last decade leading to a variety of different protocols, a massive gain in throughput, sensitivity and significant reduction in cost-per-cell compared to its early stages.
As a consequence, scRNA-seq has a need for tailored bioinformatic software solutions able to tackle the new challenges. These include protocol-specific processing of barcodes and unique molecular identifier (UMI) sequences for up to hundreds of thousands of cells in parallel, detection and characterisation of cellular clusters and appropriate visualizations.
During my thesis, I'm developing WASP - a web-accessible scRNA-seq analysis platform. WASP covers a complete workflow for data based on the ddSeq protocol, from raw reads to cell clustering, differential gene expression detection and visualization. Due to its modular design, the software can easily be employed for data from other protocols. To perform on-premise analysis of sensitive data, WASP can be employed using Docker, Conda or simply as a standalone version for Windows-based systems. Furthermore, users can interactively change parameters during the analysis workflow and download publication-ready visualizations.
PhD project of Tobias Zimmermann
Petra: A new R package for epigenome and transcriptome analysis within the Bioconductor platform.
Analysis of NGS data comes along with the requirements for computational infrastructure that allows the execution of relevant analysis tools without expert programming knowledge. We want to provide an R package, Petra, for epigenome and transcriptome analysis within the Bioconductor platform.
Petra will be accessible to the community to enable researchers to perform standard ChIP-seq, ATAC-seq, and RNA-seq analysis. In addition to basic workflows for the execution of principal analysis steps like differential expression/binding analysis and visualizations for exploratory analysis, we would like to integrate more specific and complex functionality dedicated to questions resulting from the combined analysis. For this purpose, we have implemented a super-enhancer detection algorithm based on peak position data. By combining different peak annotation approaches, a new peak to gene association algorithm has been designed. Additionally, functionality for the visualization of genomic data like browser snapshots, motif analysis, or correlation heat maps has been implemented.
The integration of transcriptome and epigenome data analysis will help put forward and test new hypotheses more efficiently. The R package Petra allows maximum flexibility to add new features. It provides a combined analysis of epigenome and transcriptome data - differential bound transcription factors and differentially expressed genes will be identified.
PhD project of Patrick Blumenkamp
The yearly increasing citations of DESeq2, edgeR, and limma (an increase of 535 % from 2015 to 2018) show that differential gene expression (DGE) analyses are still on an emerging path. The vast amount of data generated by current sequencing instruments underpins the need for automated and reproducible analysis pipelines.
Thus, we develop a two-component software for analyzing and visualizing RNA-Seq data focusing on DGE analyses. The first part is a modularized Snakemake pipeline generator consisting of quality control, preprocessing, mapping, and in-depth analysis modules, called Curare. The pipelines are built for high-throughput analyses and can be executed on local machines as well as on high-performance compute clusters. Each pipeline is entirely reproducible, and the existing collection of modules, which are customizable and extendable, increases the flexibility of the pipeline generation. The second component is a tool for visualizing DGE results. With the Gene Expression Visualizer (GenExVis), DGE results can be interactively analyzed, and numerous charts can be created. All charts can be saved in common image file formats for usage in presentations and publications. Both components combined create an environment that supports the full process of data analysis from the initial handling of RNA-seq raw data to the final DGE analyses and result visualization.
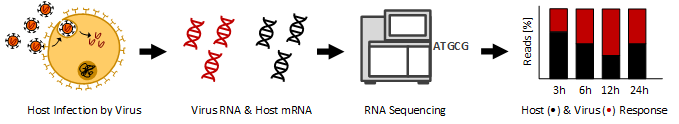
PhD project of Nina Hofmann: Integrative and comparative analysis of virus-host interactions
Virus infections remain a major threat to human health. Virus-host interactions are often not fully understood resulting in a lack of available treatment and vaccination in many cases. RNA viruses are of particular interest, because their replication machinery introduces a high number of nucleotide substitutions. This leads to high variability among the virus genomes which is an essential factor to adapt to changing environmental conditions or to new hosts.
In my thesis I analyze high-throughput RNA-Seq data taken from human pathogenic RNA viruses from different families covering the respiratory viruses human CoV-229E, MERS-CoV, a highly pathogenic H5N1 IV, a seasonal H1N1 IV and RSV, the hemorrhagic fever causing viruses Ebola virus (EBOV), Marburg virus (MARV), Lassa virus (LASV) and Rift Valley fever virus (RVFV), as well as Nipah virus (NIV), Sandfly fever Sicilian virus (SFSV), and hepatitis C virus (HCV). For this purpose, I am developing a bioinformatics pipeline that is adjusted to evaluate transcriptome changes of virus-host interactions after infection with RNA viruses over time. The RNA-Seq pipeline provides an automated workflow for the joint evaluation of host transcriptome and viral genome data.