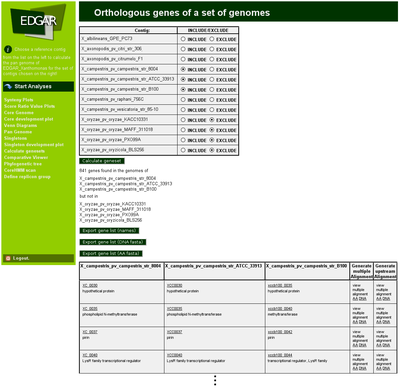
Calculate genesets
The geneset calculation allows to create complex genomic subsets like the subsets of Venn diagrams for more than 5 genomes.
In the upper part of the feature page a table shows all genomes stored in the EDGAR project and a set of select buttons with the options “INCLUDE” and “EXCLUDE” for every genome.The gene set is calculated such that there has to be a set of orthologous genes in all included genomes while there must not be any ortholog to one of the excluded genomes. Genomes where neither “INCLUDE” nor “EXCLUDE” is checked are ignored.
In the notation of set theory that means if two genomes A and B are included and a third genomes C is excluded, the subset A ∩ B \ C is calculated.