Venn diagrams
Venn diagram as implemented in EDGAR show the number of genes for all possible logical combinations of a collection of up to five genomes.
In the web interface the user can select up to five genomes or sets from the selection list. EDGAR will create a Venn diagram for the selection. The following image shows two Venn diagrams created with the EDGAR web interface:
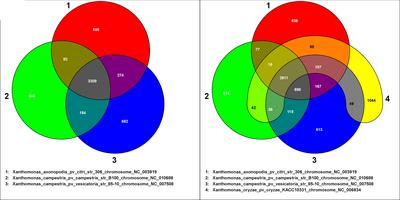
On the left side a Venn diagram of three Xanthomonas genomes is shown that illustrates the number of singleton genes (red, green, and blue ares) and the core genome (centered gray shape), but also the intersections in mixed colors. On the right side a Venn diagram of 4th order is shown resulting from the addition of another Xanthomonas strain. Again the core genome and the singleton genes appear in their respective areas, but all possible subsets of two or three genomes can be observed, too. Each included genome has one basic color that can be seen in the area representing the singletons, all other areas are colored with the combination color of the contributing genomes’ colors. The numbers in the single areas are links that forward the user to a table listing the genes of the respective subset.
EDGAR limits the order of Venn diagrams to five although Venn diagrams for six and more genomes are possible. But as the number of regions within a Venn diagram of nth order is 2n − 1 this results in too many and too small areas in the graphical representation and thus becomes too unclear to be reasonable.
To allow the user genome comparisons of higher order EDGAR provides an interface to calculate any possible intersection set of any arbitrary number of genomes in the "Calculate geneset" interface.
