Platon - identification and characterization of bacterial plasmid contigs
Platon detects plasmid contigs within bacterial draft genomes from WGS short-read assemblies. Therefore, Platon analyzes the natural distribution biases of certain protein coding genes between chromosomes and plasmids. This analysis is complemented by comprehensive contig characterizations upon which several heuristics are applied.
Workflow
Platon conducts three analysis steps:
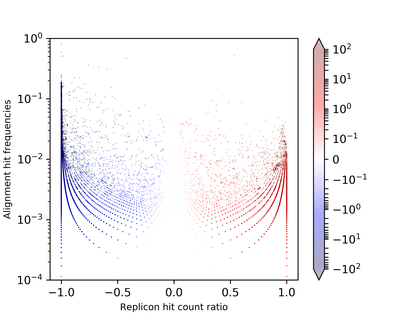
- It predicts and searches coding sequences against a custom and pre-computed database comprising marker protein sequences (MPS) and related replicon distribution scores (RDS). These scores express the empirically measured frequency biases of protein sequence distributions between plasmids and chromosomes pre-computed on complete NCBI RefSeq replicons. Platon calculates the mean RDS for each contig and either classifies them as chromosome if the RDS is below a sensitivity cutoff determined to 95% sensitivity or as plasmid if the RDS is above a specificity cutoff determined to 99.9% specificity. Exact values for these thresholds have been computed based on Monte Carlo simulations of artifical replicon fragments created from complete RefSeq chromosome and plasmid sequences.
- Contigs passing the sensitivity filter get comprehensivley characterized. Hereby, Platon tries to circularize the contig sequences, searches for rRNA, replication, mobilization and conjugation genes, oriT sequences, incompatibility group DNA probes and finally performs a BLAST+ search against the NCBI plasmid database.
- Finally, to increase the overall sensitivity, Platon classifies all remaining contigs based on the gathered information by several heuristics.
Availability & further information
- Publication in Microbial Genomics: https://doi.org/10.1099/mgen.0.000398
- All information is available at GitHub: https://github.com/oschwengers/platon
- The software is available via BioConda: https://bioconda.github.io/recipes/platon/README.html
- The mandatory database is publicly hosted at Zenodo:
