Research activities
Research activities of our group are mainly focused on the development of innovative bioinformatics software tools. The design and implementation of novel features is driven by our close collaboration with partners working in various fields of research. Within many projects, experimental data are produced by applying high-throughput methods and it is therefore one of our major goals to simplify and automate the subsequent steps for a systematic data processing and analysis as far as possible.
For the development of larger software platforms we follow the design principle to build specialized tools for separate scopes that can be interlinked to integrate the various data resources. Technically, we rely on the hardware infrastructure of the Bioinformatics Core Facility (BCF) that provides the necessary storage and compute capacities. An overview of our software platform is depicted in the figure below.
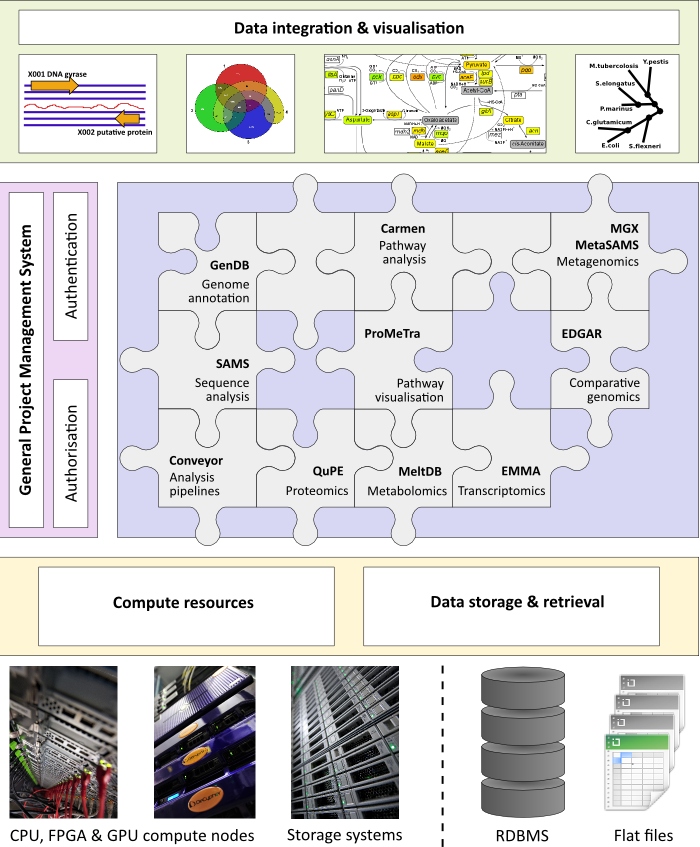
Applications developed in the field of genomics comprise the maintenance and annotation of complete genomes as well as the evaluation of other sequence data employing the GenDB software. The GenDB system was successfully used for the automatic and manual annotation of more than 150 microbial genomes. Based on the genome annotation data provided by GenDB, the CARMEN application can perform a metabolic reconstruction. CARMEN provides two main applications: the generation of de novo models based on KEGG database information and the creation of template based SBML models for comparative genomics. An organism-specific network model of individual or large-scale metabolic pathways can be reconstructed based on genome annotation data that can be obtained from GenDB or alternatively loaded from NCBI GenBank files.
In the field of transcriptomics, the ReadXplorer software is provided as a software platform for the evaluation of data resulting from genome-wide RNA-Seq studies.
