Define Metacontig / Replicon group
EDGAR features the defintion of "Metacontigs" or replicon groups for higher level comparisons.
Basically, EDGAR provides comparisons of single contigs or complete organisms, but sometimes a higher complexity is needed. For example, a level of abstraction between organism and contig is needed if someone wants to compare the gene content of all plasmids of one organism to the genes of all plasmids of another organism. For such cases EDGAR allows users to create groups of contigs of their choice. Genes of the selected replicons can be used as one group in all comparison features of EDGAR, subsequently.
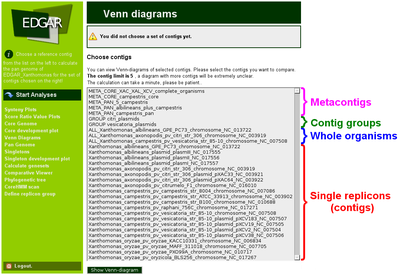
This groups work fine if they are created of contigs within one organism, but if contigs from different organisms are grouped together the redundant orthologs within the group act as artificial paralogs and prevent a reasonable analysis. Such comparisons of contig sets from different organisms or even of sets of complete organisms can be crucial to answer biological question, e.g, if a researcher wants to compare a set of pathogenic bacteria to a set of non-pathogenic bacteria. Thus, an abstraction level above organism is needed to store non-redundant sets of genes for a group of contigs or organisms. In EDGAR this is realized via so-called metacontigs. A user can define metacontigs via the web interface. First, he has to select organisms and contigs that should be grouped together with a reference genome. Second, a method to remove redundant genes has to be selected, either the pan genome or the core genome calculation. The respective genomic subset will be calculated for the contigs and organisms and a non-redundant set of genes is extracted by using one representative gene of each ortholog set from the result, preferring the gene from the reference. This non-redundant set forms the metacontig and is stored in the EDGAR database. The two user defined groups have their respective menu item in the EDGAR web interface. Contig groups have just the prefix “GROUP”, while metacontigs have a prefix “META” followed by a second prefix “PAN” or “CORE” to indicate what type of metacontig was created. A typical selection list for the calculation of a Venn diagram with all four types of menu items is shown in the following figure figure.