Overview
|
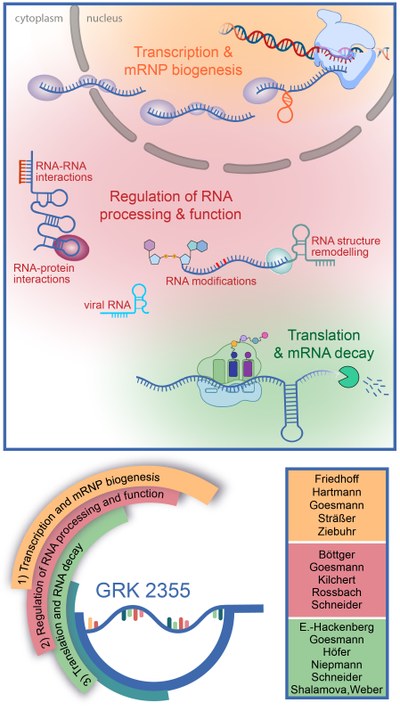
The RTG 2355 (GRK 2355) is a DFG-funded Research Training Group that brings together experts on different aspects of RNA biology to jointly address RNA-based gene-regulatory mechanisms in a broad range of model organisms. Now in its second funding period (2023 - 2027), the RTG comprises 13 groups from the Justus Liebig University (JLU) Giessen, the Max Planck Institute for Heart and Lung Research in Bad Nauheim, the Philipps University Marburg, and the Max Planck Institute for Terrestrial Microbiology in Marburg. Speaker of the RTG is Prof. Dr. Katja Sträßer, head of the Institute of Biochemistry of the Biology and Chemistry faculty of the JLU.
RNA is a central molecule of gene expression that is not only a target of regulation, but also a regulatory agent itself, and we are only beginning to appreciate how versatile the hitherto identified noncoding RNAs are used in cells: as scaffolds and recruiting platforms, protein and RNA sponges, allosteric regulators of protein activity, or molecular sensors. Functioning of these regulatory RNAs can in turn be regulated by mechanisms affecting base-pairing abilities and RNA folding such as RNA editing, base modification or RNA structure remodelling. Within our Research Training Group, we aim to establish how these multiple RNA-based mechanisms contribute to the plasticity of gene-regulatory networks. As regulation of gene expression underpins all cellular processes, this research has the potential to improve our understanding of many topics of medical relevance, such as bacterial biofilm formation, diet-induced changes to metabolism, virus infection, and disease.
Due to the large number of topics and model organisms studied within our group, and the resulting diversity in seminars within the RTG and by invited speakers, during retreats and symposia, the RTG's doctoral students gain a thorough background in RNA biology; day-to-day research on the candidates' thesis projects, internal lab rotations, workshops and short-term stays in laboratories abroad convey practical experience in state-of-the-art RNA technologies. Our qualification programme thus provides an internationally competitive research-oriented education for our dotoral students. Complemented by transferable skills courses, coaching of student assistants and peers ("learning-by-teaching") and inclusion in the organization of seminars and events ("learning-by-doing"), it furthermore enables them to address future scientific challenges in all kinds of environments, be it academia, industry, science management, publishing, patent agencies, regulatory authorities, legal affairs, or politics. |
|