RTG 2355
Overview
|
The RTG 2355 (GRK 2355) is a DFG-funded Research Training Group that brings together experts on different aspects of RNA biology to jointly address RNA-based gene-regulatory mechanisms in a broad range of model organisms. Now in its second funding period (2023 - 2027), the RTG comprises 13 groups from the Justus Liebig University (JLU) Giessen, the Max Planck Institute for Heart and Lung Research in Bad Nauheim, the Philipps University Marburg, and the Max Planck Institute for Terrestrial Microbiology in Marburg. Speaker of the RTG is Prof. Dr. Katja Sträßer, head of the Institute of Biochemistry of the Biology and Chemistry faculty of the JLU.
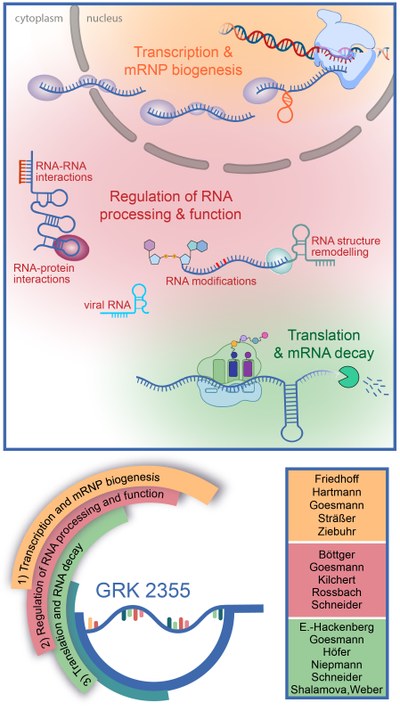
RNA is a central molecule of gene expression that is not only a target of regulation, but also a regulatory agent itself, and we are only beginning to appreciate how versatile the hitherto identified noncoding RNAs are used in cells: as scaffolds and recruiting platforms, protein and RNA sponges, allosteric regulators of protein activity, or molecular sensors. Functioning of these regulatory RNAs can in turn be regulated by mechanisms affecting base-pairing abilities and RNA folding such as RNA editing, base modification or RNA structure remodelling. Within our Research Training Group, we aim to establish how these multiple RNA-based mechanisms contribute to the plasticity of gene-regulatory networks. As regulation of gene expression underpins all cellular processes, this research has the potential to improve our understanding of many topics of medical relevance, such as bacterial biofilm formation, diet-induced changes to metabolism, virus infection, and disease.
Due to the large number of topics and model organisms studied within our group, and the resulting diversity in seminars within the RTG and by invited speakers, during retreats and symposia, the RTG's doctoral students gain a thorough background in RNA biology; day-to-day research on the candidates' thesis projects, internal lab rotations, workshops and short-term stays in laboratories abroad convey practical experience in state-of-the-art RNA technologies. Our qualification programme thus provides an internationally competitive research-oriented education for our dotoral students. Complemented by transferable skills courses, coaching of student assistants and peers ("learning-by-teaching") and inclusion in the organization of seminars and events ("learning-by-doing"), it furthermore enables them to address future scientific challenges in all kinds of environments, be it academia, industry, science management, publishing, patent agencies, regulatory authorities, legal affairs, or politics. |
|
Overview
|
The RTG 2355 (GRK 2355) is a DFG-funded Research Training Group that brings together experts on different aspects of RNA biology to jointly address RNA-based gene-regulatory mechanisms in a broad range of model organisms. Now in its second funding period (2023 - 2027), the RTG comprises 13 groups from the Justus Liebig University (JLU) Giessen, the Max Planck Institute for Heart and Lung Research in Bad Nauheim, the Philipps University Marburg, and the Max Planck Institute for Terrestrial Microbiology in Marburg. Speaker of the RTG is Prof. Dr. Katja Sträßer, head of the Institute of Biochemistry of the Biology and Chemistry faculty of the JLU.
RNA is a central molecule of gene expression that is not only a target of regulation, but also a regulatory agent itself, and we are only beginning to appreciate how versatile the hitherto identified noncoding RNAs are used in cells: as scaffolds and recruiting platforms, protein and RNA sponges, allosteric regulators of protein activity, or molecular sensors. Functioning of these regulatory RNAs can in turn be regulated by mechanisms affecting base-pairing abilities and RNA folding such as RNA editing, base modification or RNA structure remodelling. Within our Research Training Group, we aim to establish how these multiple RNA-based mechanisms contribute to the plasticity of gene-regulatory networks. As regulation of gene expression underpins all cellular processes, this research has the potential to improve our understanding of many topics of medical relevance, such as bacterial biofilm formation, diet-induced changes to metabolism, virus infection, and disease.
Due to the large number of topics and model organisms studied within our group, and the resulting diversity in seminars within the RTG and by invited speakers, during retreats and symposia, the RTG's doctoral students gain a thorough background in RNA biology; day-to-day research on the candidates' thesis projects, internal lab rotations, workshops and short-term stays in laboratories abroad convey practical experience in state-of-the-art RNA technologies. Our qualification programme thus provides an internationally competitive research-oriented education for our dotoral students. Complemented by transferable skills courses, coaching of student assistants and peers ("learning-by-teaching") and inclusion in the organization of seminars and events ("learning-by-doing"), it furthermore enables them to address future scientific challenges in all kinds of environments, be it academia, industry, science management, publishing, patent agencies, regulatory authorities, legal affairs, or politics. |
|
Organization & Contact
 |
|
Speaker
Prof. Dr. Katja Sträßer Justus-Liebig-Universität Giessen
Institut für Biochemie
Phone: +49-641-99-35400
Email: Katja.Straesser
|
 |
|
Vice-Speaker
Dr. Cornelia Kilchert Justus-Liebig-Universität Giessen
Institut für Biochemie
Phone: +49-641-99-35405
Email: Cornelia.Kilchert
|
 |
CoordinatorDr. Vera Bettenworth Justus-Liebig-Universität Giessen
Institut für Biochemie
Heinrich-Buff-Ring 17, Room B 040 Phone: +49-641-99-35405
Email: Vera.Bettenworth
|
Steering CommitteeThe steering committee is responsible for all scientific, financial, and organisational issues. It consists of the speaker, the vice-speaker, two further PIs and two doctoral student representatives: |
 |
|
responsible for the qualification programme
Apl. Prof. Dr. Elena Evguenieva-Hackenberg Justus-Liebig-Universität Giessen
Institut für Mikrobiologie
|
 |
|
responsible for career advancement measures
Dr. Katharina Höfer Max-Planck-Institut für terrestrische Mikrobiologie Marburg
Bacterial Epitranscriptomics
|
 |
|
Ph.D. representativeYannic Noe (AG Rossbach) |
 |
|
Ph.D. representativeHelene Keuthen (AG Höfer) Max-Planck-Institut für terrestrische Mikrobiologie Marburg
Bacterial Epitranscriptomics
|
Upcoming Events

Thursday, 18 April 2024, 13:00 - 18:00
Speakers: Jessica Fedrau, Yannic Noe, Fabienne Becker, Daniel Bauer, Palina Kot & Jennifer Kothe
Heinrich-Buff-Ring 26, Gießen

Monday, 6 May 2024, at 16:00
Prof. Dr. Achim Aigner, Leipzig University
Heinrich-Buff-Ring 19, Gießen
Host: Oliver Rossbach, Institute of Biochemistry, FB08, JLU

Thursday, 23 May 2024, at 16:00
Dr. Sina Wittmann, Institute of Molecular Biology, Mainz
Heinrich-Buff-Ring 26, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Thursday, 6 June 2024, at 16:00
Dr. Markus Höpfler, Medical Research Council Laboratory of Molecular Biology, Cambridge
Heinrich-Buff-Ring 26, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Tuesday, 9 July 2024, 13:00 - 18:00
Save the date!

Monday, 15 July - Wednesday, 17 July 2024
Location: Martin-Niemöller-Haus, Schmitten, Taunus

Thursday, 5 September 2024, at 16:00
Prof. Dr. Stefan Hüttelmaier, Martin-Luther-Universität Halle-Wittenberg
Location: tba
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Thursday, 10 October 2024, 13:00 - 18:00
Save the date!

Thursday, 31 October 2024, at 16:00
Dr. Andreas Mayer, Max Planck Institute for Molecular Genetics, Berlin
Heinrich-Buff-Ring 26, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Thursday, 14 November 2024, at 16:00
Prof. Dr. Petra Dersch, Universität Münster
Location: tba
Host: Elena Evguenieva-Hackenberg, Institute of Microbiology and Molecular Biology, FB08, JLU

Tuesday, 26 November 2024, at 16:00
Prof. Dr. Chase Beisel, Helmholtz Institute for RNA-based Infection Research, Würzburg
Location: tba
Host: Katharina Höfer, Max Planck Institute for Terrestrial Microbiology, Marburg
Past Events

Wednesday, 10 April - Friday, 12 April 2024
Dynamics in chromatin organization and RNA regulation: adaptation, infection - and beyond
The symposium is a joint venture with the EU-funded Marie Curie Innovative Training Network "Cell2Cell - role of cellular heterogeneity in chromatin structures in pathogen infections" and brings together international speakers and local experts in chromatin organization, RNA regulation, and pathogen biology. Keynote speakers are former FMI Basel Director Susan Gasser (Swiss Institute for Experimental Cancer Research, Lausanne), Schraga Schwartz (Weizmann Institute of Science, Rehovot), and Bas van Steensel (Netherlands Cancer Institute, Amsterdam). A welcome reception will take place on day 1, followed by a poster session on day 2, offering ample opportunity for scientific interactions. For a detailed programme and to register, please check the symposium website. Registration will be open till Friday, 15 March.
Ludwigstr. 23, 35390 Giessen

Tuesday, 9 April, 10:00 - 16:00
What Comes Next
This workshop aims at helping you identify and begin to prepare for your next career step. Moreover, you will further develop your capacity to take responsibility and acquire basic tools to support you in day-to-day supervision.
Heinrich-Buff-Ring 17, Gießen
Trainers: Conor John Fitzsimons & Petra Pandur, Leadership Sculptor

Monday, 8 April, 10:00 - 17:30
Introduction to Project Management
This workshop will introduce you to basic project management tools, help you identify and overcome obstacles to your scientific maturation, and explore and develop some new research ideas.
Heinrich-Buff-Ring 17, Gießen
Trainer: Conor John Fitzsimons, Leadership Sculptor

Monday, 18 March & Tuesday, 19 March 2024, both 9:00 - 16:00
Introduction to Galaxy analyses & Reference-based RNA-Seq data analysis
Galaxy is a worldwide open source project with the European Galaxy Server being the biggest instance in Europe with more than 85,000 users. The Freiburg Galaxy Team is hosting this server in Freiburg. Through Galaxy as a gateway, we are offering free access to a huge computational cloud infrastructure, databases and 3,200 bioinformatics tools which can be used by a graphical user interface instead of command-line. There is no need for programming or informatics skills - you just need a web browser (e.g. chrome or firefox). We will have demonstrations and work together on detailed E-learning step-by-step-instructions of the Galaxy Training Material.
Heinrich-Buff-Ring 58, JLU
Trainers: Dr. Anika Erxleben, Universität Freiburg, European Galaxy Team; Jochen Blom, Sven Griep & Oliver Rupp, Justus Liebig University Giessen, de.NBI BIGI; Fabienne Thelen, Justus Liebig University Giessen, RTG 2355

Thursday, 14 March 2024, at 16:00
Prof. Dr. Mark Helm, Johannes Gutenberg University Mainz
Heinrich-Buff-Ring 26, Gießen
Host: Maximilian Staps, Max Planck Institute for Heart and Lung Research, Bad Nauheim

Monday, 4 March 2024, at 16:15
Prof. Dr. Tobias Jakobi, University of Arizona, Phoenix
"Computational detection and analysis of circular RNAs in the heart"
Heinrich-Buff-Ring 19, Gießen
Host: Jochen Blom, Bioinformatics & Systems Biology Group, FB08, JLU

Tuesday, 27 February 2024, at 16:00
Prof. Dr. Danny Nedialkova, Max Planck Institute of Biochemistry, Martinsried & Technical University of Munich
"Translational control in development and differentiation"
Heinrich-Buff-Ring 19, Gießen
Host: Cornelia Kilchert, Institute of Biochemistry, FB08, JLU

Tuesday, 20 February 2024, at 15:00
Prof. Dr. Federica Accornero, Brown University, Rhode Island
"RNA methylation in the regulation of muscle homeostasis"
Heinrich-Buff-Ring 19, Gießen
Host: Maximilian Staps, Max Planck Institute for Heart and Lung Research, Bad Nauheim

Wednesday, 14 February & Thursday, 15 February 2024
Presenting for success - Designing and delivering scientific talks
The workshop starts with a brief recap of some principles that all presenters need to bear in mind for addressing any audience. Drawing on these, we will then dig into the practical consequences for designing a presentation and structuring its information content for maximum effect. Even more practically (!) we will consider things such as where to stand, how to speak and move, and the physical relationship between presenter and audience. Interspersed throughout the workshop we will have the pleasure of being the audience for presentations made by you, the participants. Please, therefore, make a brief presentation of your work in powerpoint or similar, and be prepared to present it at the workshop without exceeding 5 minutes.
Timeline: day 1 & day 2: 10:00 - 16:00 with 1 hour lunch break
Heinrich-Buff-Ring 17, Gießen
Coach: Andrew Moore

Thursday, 1 February 2024, at 16:00
Dr. Sonja Lorenz, Max Planck Institute for Multidisciplinary Sciences, Göttingen
“Structural mechanism of the ubiquitin ligase HACE1”
Heinrich-Buff-Ring 19, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Tuesday, 23 January 2024, 14:15 - 18:00
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU
Chair: Fabienne Becker, JLU
Heinrich-Buff-Ring 26, Gießen

Tuesday, 12 December 2023, at 16:00
Prof. Dr. Markus Landthaler, Max Delbrück Center Berlin
"Posttranscriptional regulation in cellular space and time"
Heinrich-Buff-Ring 19, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Monday, 4 December & Tuesday, 5 December 2023
Scientific presentations – Concise content, storytelling & slide decks
Within this 1,5 day-workshop, participants will learn to deliver high quality scientific talks with ease. Key elements are (1) to learn how to simplify complex content without losing precision, (2) to incorporate storytelling elements to communicate complex ideas effectively and (3) to create visually appealing slide decks that enhance comprehension. To this end, the training combines short, digestible learning snippets with interactive practice sessions, ensuring that participants have ample opportunity to apply their newly acquired knowledge in a supportive environment providing valuable feedback.
Pizza & personalized tips on day 1 for those who are willing to stay late.
Heinrich-Buff-Ring 26, Gießen
Coach: Simon Klug
The follow-up workshop Scientific presentations – Pitching, key messages & stage presence – is scheduled for the first week of February 2024.

Thursday, 30 November 2023, at 16:00
Prof. Dr. Beatrix Süß, Technische Universität Darmstadt
"Next-level riboswitch development - Implementation of Capture-SELEX allows fast and easy identification of synthetic riboswitches"
Heinrich-Buff-Ring 26, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Tuesday, 14 November 2023, at 16:00
Prof. Dr. Maria Hondele, Biozentrum Basel
Heinrich-Buff-Ring 19, Gießen
Host: Cornelia Kilchert, Institute of Biochemistry, FB08, JLU

Thursday, 9 November & Friday, 10 November 2023
Think before you write - Properly representing your research in the online age and understanding publishing
The workshop starts with insights into the cognitive psychology of readers, from which we will draw very practical consequences for writing and structuring our manuscripts. There will be short exercises and longer assignments for participants throughout the workshop, and all participants should submit a title and abstract of their work before the course: these will be worked on during the workshop with input from the workshop coach.
Timeline: day 1 & day 2: 10:00 - 16:00 with 1 hour lunch break
Heinrich-Buff-Ring 17, Gießen
Coach: Andrew Moore

Thursday, 2 November 2023, at 16:00
Prof. Dr. Utz Fischer, Universität Würzburg
"Making mRNA in the wrong compartment: how poxviruses express their genes in the cytoplasm of a host"
Heinrich-Buff-Ring 19, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Tuesday, 24 October 2023, 16:00 - 18:00
Unconscious bias II - online follow-up
Coach: Anna Schwark & Maria Prahl, Working Between Cultures

Thursday, 19 October 2023, at 16:00
Dr. Björn Rotter, GenXPro, Frankfurt
"RNA-based information for therapeutic decision support for cancer"
Heinrich-Buff-Ring 26, Gießen
Host: Jochen Blom, Bioinformatics & Systems Biology group, JLU

Tuesday, 10 October 2023, 13:00 - 18:00
Unconscious bias I
Navigating the academic journey with clarity & empathy: unconscious bias awareness for scientists
In this mandatory half-day workshop, we will uncover unconscious biases in our environment and in ourselves, understand the origins of these biases and recognize how they affect our day-to-day lives on all levels. We will experience a change of perspective and learn how to navigate stereotypes to shape an (academic) environment based on mutual respect and empathy in which everybody's ideas can be heard.
Heinrich-Buff-Ring 26, Gießen
An online follow-up workshop is scheduled for Tuesday, 24 October, 16:00 - 18:00.
Coach: Anna Schwark, Working Between Culture

Wednesday, 4 October - Saturday, 7 October 2023
12th Meeting of the GBM Study Section "RNA Biochemistry" & Workshop "Advanced RNA Sequencing"
Tagungs- und Gästehaus CJD Castell, Graurheindorfer Str. 149, Bonn
Keynote speakers: Danny Nedialkova, MPI of Biochemistry Martinsried & TU Munich & Maria Hondele, Biozentrum, University of Basel
Click for Programme & further information

Thursday, 28 September 2023, 12:30 - 18:00
RMU-RNA Salon "Genomics approaches in RNA biology" - 11. Minisymposium "RNA virus research"
Heinrich-Buff-Ring 19, Giessen
Keynote speakers: Susanne Herold, UKGM Gissen & John Ziebuhr, JLU Giessen
Click for Flyer & Registration

Wednesday, 20 September & Thursday, 21 September 2023
16th GGL Annual Conference
Heinrich-Buff-Ring 14, Giessen
GGL doctoral researchers of all ten research sections will contribute to the programme and present their results. Each session will be opened by a guest speaker who is an expert in the specific field. Prizes will be awarded in the categories best talk and best photograph.
Click for Conference Website & Programme

Thursday, 20 July 2023, at 16:00
Dr. Gabrijela Dumbovic, Goethe-Universität Frankfurt
"RNA localization controls RNA function"
Heinrich-Buff-Ring 19, Gießen
Host: Theresa Gerhardt, Max Planck Institute for Heart and Lung Research, Bad Nauheim

Wednesday, 5 July - Friday, 7 July 2023
Joint retreat with RTG 2344
Location: Bad Dürkheim, Rheinland-Pfalz

Wednesday, 28 June - Friday, 30 June 2023
Programming with R
Oliver Rupp, Sven Griep, Patrick Barth, Fabienne Thelen, Karina Brinkrolf, Bioinformatics & Systems Biology group, JLU
This workshop will provide a general introduction to programming with R, data visualization with ggplot2 and statistical tests, using RNA-seq data as an example. Previous knowledge in R is not required.

Tuesday, 20 June 2023, at 16:00
Prof. Dr. Michaela Müller-McNicoll, Goethe-Universität Frankfurt
"The role of subcellular architecture in the regulation of gene expression during stress"
Heinrich-Buff-Ring 26, Gießen
Host: Ebru Aydin, Institute of Biochemistry, FB08, JLU

Thursday, 15 June 2023, 13:00 - 18:00
Speakers: Fabienne Becker, Alexander Goesmann, Fabian Dross, Cornelia Kilchert, Oliver Rossbach & Kiriaki Kouti
Chairs: Maximilian Staps, Yannic Noe & Ebru Aydin
Heinrich-Buff-Ring 19, Gießen

Monday, 5 June 2023, at 16:00
Prof. Dr. Julia Weigand, Philipps-Universität Marburg
"mRNA structure shapes human and viral gene regulation"
Heinrich-Buff-Ring 19, Gießen
Host: Oliver Rossbach, Institute of Biochemistry, FB08, JLU

Wednesday, 31 May 2023
Introduction to LaTex
Dr. Jochen Blom, Bioinformatics & Systems Biology group, JLU
Introduction to the software including hands-on training.

Tuesday, 16 May 2023, at 16:00
Dr. Julian König, Institute of Molecular Biology, Mainz
"RNA stability controlled by m6A methylation mediates X-to-autosome dosage compensation in mammals"
Heinrich-Buff-Ring 26, Gießen
Host: Maximilian Staps, Max Planck Institute for Heart and Lung Research, Bad Nauheim

Tuesday, 2 May 2023, at 16:00
Prof. Dr. Harald Schwalbe, Goethe-Universität Frankfurt
"Understanding the mechanism of translational and transcriptional riboswitches"
Heinrich-Buff-Ring 26, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Thursday, 27 April 2023, at 13:00
Speakers: Ebru Aydin, Ahmad Altoun, Katja Sträßer, Sophie Stebel, Maximilian Staps & Lyudmila Shalamova
Chair: Jennifer Kothe, Theresa Gerhardt & Daniel Bauer
Heinrich-Buff-Ring 19, Gießen

Monday, 27 March 2023, at 16:00
Prof. Dr. Steven West, University of Exeter
"A good beginning makes for a great ending: mechanisms of promoter-proximal transcriptional termination"
Heinrich-Buff-Ring 19, Gießen
Host: Theresa Gerhardt, Max Planck Institute for Heart and Lung Research, Bad Nauheim

Thursday, 16 March 2023, at 16:00
Dr. Marco Preußner, Freie Universität Berlin
"Mechanisms and functional consequences of regulated 3' splice site selection after step 1 of splicing"
Heinrich-Buff-Ring 19, Gießen
Host: Oliver Rossbach, Institute of Biochemistry, FB08, JLU

Tuesday, 7 March 2023, at 16:00
Speakers: Theresa Gerhardt, Katharina Höfer & Peter Friedhoff
Chair: Kai Wallerang
Online (BigBlueButton)

Thursday, 2 March 2023, at 16:00
Prof. Dr. Anna-Lena Steckelberg, Columbia University, New York
"Viral RNA structures as master manipulators of cellular machinery"
Heinrich-Buff-Ring 19, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

Thursday, 23 February 2023, at 16:00
Prof. Dr. Argyris Papantonis, Georg-August-Universität Göttingen
"RNA and speckles shaping the senescent nucleus"
Heinrich-Buff-Ring 19, Gießen
Host: Ebru Aydin, Institute of Biochemistry, FB08, JLU

Thursday, 2 February 2023, at 16:00
Prof. Dr. Stefan Janssen, Justus-Liebig-Universität Giessen
"RNA secondary structure prediction"
Heinrich-Buff-Ring 19, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU

RTG2355/2 kick-off meeting
Tuesday, 24 January 2023, at 14:15
Voting of the new Steering Committee
Speakers: Patrick Barth, Elene Evguenieva-Hackenberg, Yannic Noe, Thomas Böttger
Chairs: Jacqueline Böhme & Ahmad Altoun
Heinrich-Buff-Ring 19, Gießen

Thursday, 19 January 2023, at 16:00
Prof. Dr. Petra Wendler, Universität Potsdam
"Cryo-EM resolution revolution: now seeing electrons"
Heinrich-Buff-Ring 14, Gießen
Host: Katja Sträßer, Institute of Biochemistry, FB08, JLU
News & Impressions
JLU immunologist Andreas Krueger joins with a project on miRNAs in T cell development
"Making mRNA in the wrong compartment" - Guest Lecture by Utz Fischer
Joint retreat with RTG 2344 in Bad Dürkheim, 2023
JLU bioinformatician Stefan Janssen included in the RTG
Retreat to the Silbersee, 2022
Publications
2024
The enigmatic epitranscriptome of bacteriophages: putative RNA modifications in viral infections. Curr Opin Microbiol. 12 Jan 2024, doi: 10.1016/j.mib.2023.102417
Scheuer R, Kothe J, Wähling J, Evguenieva-Hackenberg E. Analysis of sRNAs and Their mRNA Targets in Sinorhizobium meliloti: Focus on Half-Life Determination. In: Arluison, V., Valverde, C. (eds) Bacterial Regulatory RNA. Methods in Molecular Biology, vol 2741. Humana, New York, NY. doi: 10.1007/978-1-0716-3565-0_13
2023
Schoen A, Hölzer M, Müller MA, Wallerang KB, Drosten C, Marz M, Lamp B, Weber F. Functional comparisons of the virus sensor RIG-I from humans, the microbat Myotis daubentonii, and the megabat Rousettus aegyptiacus, and their response to SARS-CoV-2 infection. J Virol, 31 Oct 2023, doi: 10.1128/jvi.00205-23
Processing and decay of 6S-1 and 6S-2 RNAs in Bacillus subtilis. RNA, Oct 2023, doi: 10.1261/rna.079666.123
Hadjeras L, Heiniger B, Maaß S, Scheuer R, Gelhausen R, Azarderakhsh S, Barth-Weber S, Backofen R, Becher D, Ahrens C, Sharma C, Evguenieva-Hackenberg E. Unraveling the small proteome of the plant symbiont Sinorhizobium meliloti by ribosome profiling and proteogenomics. microLife, 10 Mar 2023, doi:10.1093/femsml/uqad012
Schott S, Scheuer R, Ermoli F, Glatter T, Evguenieva-Hackenberg E., Andreas Diepold. A ParDE toxin–antitoxin system is responsible for the maintenance of the Yersinia virulence plasmid but not for type III secretion-associated growth inhibition. Front. Cell. Infect. Microbiol., 09 May 2023, doi:10.3389/fcimb.2023.1166077
Wiegard JC, Damm K, Lechner M, Thölken C, Ngo S, Putzer H, Hartmann RK. Processing and decay of 6S-1 and 6S-2 RNAs in Bacillus subtilis. RNA, Jun 27 2023, doi: 10.1261/rna.079666.123
First Funding Period 2018-2022
| Titel | Inhaltstyp | Bild | |
|---|---|---|---|
| First Funding Period 2018-2022 | Seite | ||
| Research & qualification programme | Seite | ||
| Projects / Principal Investigators | Seite | ||
| PhD Students / Projects | Seite | ||
| Highlights from PhD Students | Seite | ||
| Publications of PhD Students | Seite | ||
| Student Seminar Series | Seite | ||
| PI Seminar Talks | Seite | ||
| Guest Lecture Series | Seite | ||
| Workshops | Seite | ||
| International Meeting (2022) | Seite | ||
| Retreat (2021) | Seite | ||
| Symposium (2019) | Seite | ||
| Retreat (2018) | Seite | ||
| Past Events | Ordner |
Fotos, Bilder, PDFs
Retreat November 2022, Ferienpark Silbersee - Photo: Ferienpark Silbersee
Elena Evguenieva-Hackenberg - Photo: Katrina Friese